Research
Aims
Already Pringsheim stated that it was not possible to distinguish species by morphology in the genus Cryptomonas, because characters proved to be unreliable and/or indistinctive (Pringsheim 1968). Cryptophytes obviously share the problems of the morphospecies concept with other protists. One aim of my research therefore is the establishment of a sound systematics with a reproducible species concept in congruence with cryptophyte phylogeny. It will facilitate testing of the resolution of DNA barcoding markers and is a prerequisite for a reliable assessment of cryptophyte diversity in biodiversity surveys (Meyer and Paulay 2005).
Another aim of my research is the elucidation of morphological and molecular character evolution. Cryptophytes e.g. invented a unique light harvesting complex. The biliproteins originate from the red algal ancestor of the complex plastids, but cryptophytes use them in a highly modified form, quite different from phycobilisomes (Glazer and Wedemayer 1995; Wilk et al. 1999). I want to understand the evolutionary history of the cryptophyte biliproteins. How did they evolve and what are their advantages over phycobilisomes?
Plastid-containing cryptophytes contain four genomes in a cell that allow for comparisons of phylogenetic trees across genomes. These comparative analyses give insight into evolution at molecular level. Changes in selective pressures also shape the DNA of an organism, i.e. may increase mutation rates and/or alter codon usage in protein-coding genes.
Methods and Research Strategy
A variety of methods help elucidating the evolution of phenotypic and molecular characters.
Cryptophytes produce at least eight different types of biliproteins (Hill and Rowan 1989; Hoef-Emden 2008).
Isolation of the biliproteins is easy, they are water-soluble and each clonal culture contains only one type of
biliprotein (Hill and Rowan 1989). Red biliprotein types (phycoerythrins) show only one peak in their respective
absorption spectra, whereas blue biliproteins (phycocyanins
have two absorption maxima.
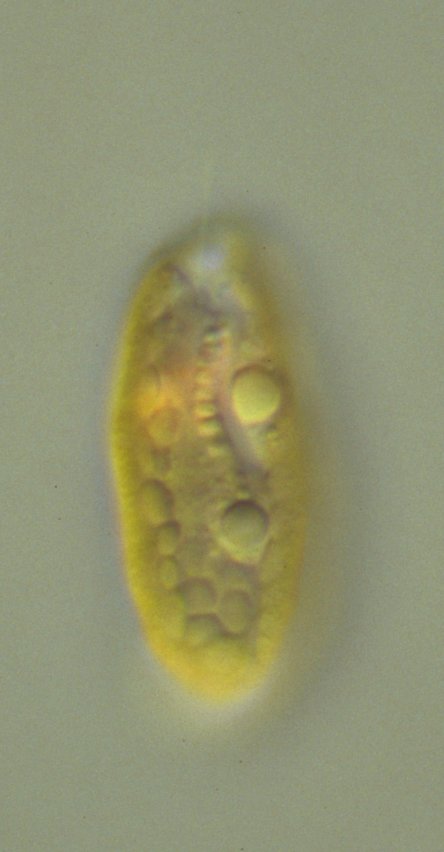
Morphological characters of live cells immobilized in ultra-low gelling agarose can be examined with
differential interference contrast (DIC) light microscopy. Periplasts, the cryptophyte-specific cell surroundings,
are isolated by adding a detergent
in a cytoskeleton-stabilizing buffer to the cells. The detergent dissolves the plasma membrane in unprotected areas of the
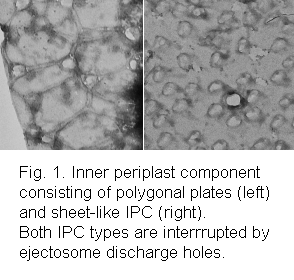
cell surface (the furrow-gullet system) and the cell contents are released leaving empty periplasts behind. Stained with
uranyl acetate, the ultrastructure of the periplasts can be examined with a transmission electron microscope (Fig. 1).
To gain insight into the evolution of phenotypic characters, the characters are mapped onto phylogenetic
trees. Resolution of the phylogenetic trees is increased by using data sets of concatenated genes of different genomes.
Prior to combining genes to a supermatrix alignment to yield higher resolved phylogenies, the single-gene phylogenies need to be compared to each other to rule out incongruences. Different from protein-coding genes, horizontal gene transfer has never been reported from ribosomal RNA genes. If discrepancies in taxon positions are found in phylogenies inferred from different ribosomal RNA genes, the most likely causes are mix-ups of strains, of DNA, of PCR products or of taxon labels. Comparisons of single-gene phylogenies provide good means for a plausibility control of ones own data. As an interesting by-result these comparisons unveil lineage-specific changes in mutation rates that may be caused by alterations in selective pressures on the cells, e.g. by a switch from a photoautotrophic or mixotrophic to a heterotrophic mode of nutrition (Hoef-Emden 2005; Hoef-Emden et al. 2005).
Observations of gametic fusions between cryptophyte cells have been reported from field samples and from culture (Wawrik 1969, 1971, 1979; Kugrens and Lee 1988). The biological species concept, thus, theoretically can be applied and at least some cryptophyte lineages seem to pass through a dimorphic life history (Hill and Wetherbee 1986; Hoef-Emden and Melkonian 2003). Since it is not yet possible to induce mating in culture, neither biological species limits can be tested nor are the details of the life history known. Thus, CBC clades are used to safely determine non-interbreedable units and to erect a reproducible species concept, that can be expanded and refined once biological species limits will be testable (Hoef-Emden 2007). To identify potential distinctive and reliable morphological characters, it is important to examine also the morphology of the cells and to add these characters together with the molecular signatures to the species diagnoses (Hoef-Emden and Melkonian 2003; Hoef-Emden 2007).
Major Results
- Evidence for dimorphism in the freshwater genus Cryptomonas: Campylomonas is an alternative morphotype of Cryptomonas; first revision of Cryptomonas (Hoef-Emden and Melkonian 2003)
- Polyphyly of heterotrophic Cryptomonas species and increased evolutionary rates in nucleomorph small subunit ribosomal RNA genes in one lineage of Cryptomonas comprising photosynthetic and heterotrophic species (Hoef-Emden 2005)
- Presence of rbcL genes with increased evolutionary rates and changed codon usage in a heterotrophic lineage of Cryptomonas (Hoef-Emden et al. 2005)
- Lack of reliable and distinctive morphological characters in pyrenoid-less taxa of the genus Cryptomonas and application of the CBC clade concept to distinguish species; second revision of Cryptomonas (Hoef-Emden 2007)
- Identification of an eighths type of biliprotein (PC577), insight into the complex evolution of blue biliproteins and demonstration of a wide-spread geographical distribution of species in the genus Chroomonas (Hoef-Emden 2008)